Spatial deconvolution without reference scRNA-seq¶
This is a tutorial on an example real Spatial Transcriptomics (ST) data (CID44971_TNBC) from Wu et al., 2021. Raw tutorial could be found in https://starfysh.readthedocs.io/en/latest/notebooks/Starfysh_tutorial_real.html
Starfysh performs cell-type deconvolution followed by various downstream analyses to discover spatial interactions in tumor microenvironment. Specifically, Starfysh looks for anchor spots (presumably with the highest compositions of one given cell type) informed by user-provided gene signatures (see example) as priors to guide the deconvolution inference, which further enables downstream analyses such as sample integration, spatial hub characterization, cell-cell interactions, etc. This tutorial focuses on the deconvolution task. Overall, Starfysh provides the following options:
At omicverse, we have made the following improvements:
Easier visualization, you can use omicverse unified visualization for scientific mapping
Reduce installation dependency errors, we optimized the automatic selection of different packages, you don’t need to install too many extra packages and cause conflicts.
Base feature:
Spot-level deconvolution with expected cell types and corresponding annotated signature gene sets (default)
He, S., Jin, Y., Nazaret, A. et al. Starfysh integrates spatial transcriptomic and histologic data to reveal heterogeneous tumor–immune hubs. Nat Biotechnol (2024). https://doi.org/10.1038/s41587-024-02173-8
import cv2
import scanpy as sc
import omicverse as ov
ov.style(font_path='Arial')
🔬 Starting plot initialization...
Using already downloaded Arial font from: /tmp/omicverse_arial.ttf
Registered as: Arial
🧬 Detecting GPU devices…
✅ NVIDIA CUDA GPUs detected: 1
• [CUDA 0] NVIDIA H100 80GB HBM3
Memory: 79.1 GB | Compute: 9.0
____ _ _ __
/ __ \____ ___ (_)___| | / /__ _____________
/ / / / __ `__ \/ / ___/ | / / _ \/ ___/ ___/ _ \
/ /_/ / / / / / / / /__ | |/ / __/ / (__ ) __/
\____/_/ /_/ /_/_/\___/ |___/\___/_/ /____/\___/
🔖 Version: 1.7.9rc1 📚 Tutorials: https://omicverse.readthedocs.io/
✅ plot_set complete.
Step 1: Prepare spatial transcriptomics (1 min)¶
Purpose: load 10x Visium (Space Ranger outputs) or similar to obtain a coordinate-aware spatial AnnData (adata_sp).
Inputs: Visium count matrix and spatial coordinates (from the
spatialfolder)Outputs:
AnnDataobject (adata_sp) with spot coordinates and countsKey points:
Ensure maximal gene overlap with the scRNA-seq reference; map gene IDs if necessary.
For multiple samples, keep batch labels explicit to support merging and visualization.
adata_sp = sc.datasets.visium_sge(sample_id="V1_Human_Lymph_Node")
adata_sp.obs['sample'] = list(adata_sp.uns['spatial'].keys())[0]
adata_sp.var_names_make_unique()
reading /scratch/users/steorra/analysis/omic_test/data/V1_Human_Lymph_Node/filtered_feature_bc_matrix.h5
(0:00:00)
Step 2: Prepare the gene sig marker¶
gene_sig means the dataframe stored the marker gene in each columns. If you don’t have it, you can calculated it using ov.space.calculate_gene_signature
# Load your scRNA-seq reference
# Example: Human lymph node reference
adata_sc = ov.datasets.sc_ref_Lymph_Node()
gene_sig=ov.space.calculate_gene_signature(
adata_sc,
clustertype='Subset', # cell type name
rank=True,
key='rank_genes_groups',
foldchange=2,
topgenenumber=20
)
gene_sig.head()
🧬 Loading SC reference data for Lymph Node
🔍 Downloading data to ./data/sc_ref_Lymph_Node.h5ad
⚠️ File ./data/sc_ref_Lymph_Node.h5ad already exists
Loading data from ./data/sc_ref_Lymph_Node.h5ad
✅ Successfully loaded: 73260 cells × 10237 genes
...get cell type marker
ranking genes
WARNING: It seems you use rank_genes_groups on the raw count data. Please logarithmize your data before calling rank_genes_groups.
finished: added to `.uns['rank_genes_groups']`
'names', sorted np.recarray to be indexed by group ids
'scores', sorted np.recarray to be indexed by group ids
'logfoldchanges', sorted np.recarray to be indexed by group ids
'pvals', sorted np.recarray to be indexed by group ids
'pvals_adj', sorted np.recarray to be indexed by group ids (0:01:00)
| B_Cycling | B_GC_DZ | B_GC_LZ | B_GC_prePB | B_IFN | B_activated | B_mem | B_naive | B_plasma | B_preGC | ... | T_CD4+_TfH | T_CD4+_TfH_GC | T_CD4+_naive | T_CD8+_CD161+ | T_CD8+_cytotoxic | T_CD8+_naive | T_TIM3+ | T_TfR | T_Treg | VSMC | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | HMGN2 | CD79B | CD22 | BIK | HLA-DPA1 | HLA-DPA1 | HLA-DPA1 | HLA-DPA1 | SDF2L1 | HLA-DPA1 | ... | TRAC | TCF7 | RPS15A | ZFP36 | CD3E | NOSIP | TMSB4X | CD4 | TRAC | TAGLN |
| 1 | DEK | TCEA1 | PRPSAP2 | PRPSAP2 | ISG15 | HVCN1 | CD52 | HVCN1 | SSR3 | CD72 | ... | CD69 | SRGN | NOSIP | SRGN | CST7 | TCF7 | LCK | SRGN | SRGN | LGALS3 |
| 2 | HMGN1 | GAPDH | HMGN1 | VPREB3 | STAT1 | CD74 | SMIM14 | CD79B | PPIB | NME2 | ... | SRGN | PASK | TCF7 | RGCC | GZMK | RGCC | SRGN | CD3E | RGCC | CST3 |
| 3 | GAPDH | SUGCT | GAPDH | HMCES | CD74 | TCL1A | CD74 | CD72 | SEC61G | CD74 | ... | CD3E | ITM2A | RPS14 | CD8A | NKG7 | DNAJB1 | CD3E | ZAP70 | DNAJB1 | LAPTM4A |
| 4 | HMGB1 | EZR | CD74 | MEF2B | IFIT3 | HLA-DPB1 | VPREB3 | CD74 | HERPUD1 | HSP90AB1 | ... | SLC2A3 | CD3E | LEF1 | CCL5 | CTSW | LEF1 | CD27 | TCRA_VDJsum | FOXP3 | IGFBP5 |
5 rows × 34 columns
gene_sig.to_csv('data/gene_sig_Lymph_Node.csv')
gene_sig=ov.pd.read_csv('data/gene_sig_Lymph_Node.csv',index_col=0)
gene_sig.head()
| B_Cycling | B_GC_DZ | B_GC_LZ | B_GC_prePB | B_IFN | B_activated | B_mem | B_naive | B_plasma | B_preGC | ... | T_CD4+_TfH | T_CD4+_TfH_GC | T_CD4+_naive | T_CD8+_CD161+ | T_CD8+_cytotoxic | T_CD8+_naive | T_TIM3+ | T_TfR | T_Treg | VSMC | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | HMGN2 | CD79B | CD22 | BIK | HLA-DPA1 | HLA-DPA1 | HLA-DPA1 | HLA-DPA1 | SDF2L1 | HLA-DPA1 | ... | TRAC | TCF7 | RPS15A | ZFP36 | CD3E | NOSIP | TMSB4X | CD4 | TRAC | TAGLN |
| 1 | DEK | TCEA1 | PRPSAP2 | PRPSAP2 | ISG15 | HVCN1 | CD52 | HVCN1 | SSR3 | CD72 | ... | CD69 | SRGN | NOSIP | SRGN | CST7 | TCF7 | LCK | SRGN | SRGN | LGALS3 |
| 2 | HMGN1 | GAPDH | HMGN1 | VPREB3 | STAT1 | CD74 | SMIM14 | CD79B | PPIB | NME2 | ... | SRGN | PASK | TCF7 | RGCC | GZMK | RGCC | SRGN | CD3E | RGCC | CST3 |
| 3 | GAPDH | SUGCT | GAPDH | HMCES | CD74 | TCL1A | CD74 | CD72 | SEC61G | CD74 | ... | CD3E | ITM2A | RPS14 | CD8A | NKG7 | DNAJB1 | CD3E | ZAP70 | DNAJB1 | LAPTM4A |
| 4 | HMGB1 | EZR | CD74 | MEF2B | IFIT3 | HLA-DPB1 | VPREB3 | CD74 | HERPUD1 | HSP90AB1 | ... | SLC2A3 | CD3E | LEF1 | CCL5 | CTSW | LEF1 | CD27 | TCRA_VDJsum | FOXP3 | IGFBP5 |
5 rows × 34 columns
Step 3: Deconvolution with starfysh¶
We perform n_repeat random restarts and select the best model with lowest loss for parameter c (inferred cell-type proportions):
Starfysh is integrated into the omicverse.space.Deconvolution class. Simply set method='starfysh'.
Key Parameters¶
n_repeats: Number of restart to run Starfysh.epochs: number of iterationsdevice: the trainning device
# Initialize the Deconvolution object
decov_obj = ov.space.Deconvolution(
adata_sp=adata_sp
)
decov_obj.preprocess_sp(
mode='pearsonr',n_svgs=3000,target_sum=1e4,
subset_genes=False,
)
🔍 [2026-01-11 23:07:13] Running preprocessing in 'cpu' mode...
Begin robust gene identification
After filtration, 25187/36601 genes are kept.
Among 25187 genes, 22411 genes are robust.
✅ Robust gene identification completed successfully.
Begin size normalization: shiftlog and HVGs selection pearson
🔍 Count Normalization:
Target sum: 10000.0
Exclude highly expressed: True
Max fraction threshold: 0.2
⚠️ Excluding 1 highly-expressed genes from normalization computation
Excluded genes: ['IGKC']
✅ Count Normalization Completed Successfully!
✓ Processed: 4,035 cells × 22,411 genes
✓ Runtime: 0.47s
🔍 Highly Variable Genes Selection (Experimental):
Method: pearson_residuals
Target genes: 3,000
Theta (overdispersion): 100
✅ Experimental HVG Selection Completed Successfully!
✓ Selected: 3,000 highly variable genes out of 22,411 total (13.4%)
✓ Results added to AnnData object:
• 'highly_variable': Boolean vector (adata.var)
• 'highly_variable_rank': Float vector (adata.var)
• 'highly_variable_nbatches': Int vector (adata.var)
• 'highly_variable_intersection': Boolean vector (adata.var)
• 'means': Float vector (adata.var)
• 'variances': Float vector (adata.var)
• 'residual_variances': Float vector (adata.var)
Time to analyze data in cpu: 3.05 seconds.
computing PCA
with n_comps=50
finished (0:00:01)
✅ Preprocessing completed successfully.
Added:
'highly_variable_features', boolean vector (adata.var)
'means', float vector (adata.var)
'variances', float vector (adata.var)
'residual_variances', float vector (adata.var)
'counts', raw counts layer (adata.layers)
End of size normalization: shiftlog and HVGs selection pearson
✓ spatial transcriptomics data is preprocessed
%time
# Run FlashDeconv deconvolution
decov_obj.deconvolution(
method='starfysh',
gene_sig=gene_sig,
starfysh_kwargs={
'n_repeats':3,
'epochs':200,
'patience':50,
'device':None,
'batch_size':32,
'alpha_mul':50,
'lr':1e-4,
'poe':False,
'verbose':True,
'n_anchors':60,
'window_size':3,
}
)
CPU times: user 3 μs, sys: 0 ns, total: 3 μs
Wall time: 7.87 μs
30 components are retained using conditional_number=40.00
Epoch[10/200], train_loss: 1964.5555, train_reconst: 1717.3358, train_u: 17.7731,train_z: 12.2491,train_c: 216.1686,train_l: 1.0289
Epoch[20/200], train_loss: 1836.5867, train_reconst: 1631.5397, train_u: 17.3562,train_z: 8.2599,train_c: 177.4633,train_l: 1.9675
Epoch[30/200], train_loss: 1803.6228, train_reconst: 1611.8570, train_u: 17.0200,train_z: 6.8935,train_c: 165.4288,train_l: 2.4234
Epoch[40/200], train_loss: 1784.1136, train_reconst: 1599.9807, train_u: 16.7433,train_z: 6.1974,train_c: 158.4979,train_l: 2.6943
Epoch[50/200], train_loss: 1771.6689, train_reconst: 1591.6377, train_u: 16.5159,train_z: 5.8556,train_c: 154.8143,train_l: 2.8453
Epoch[60/200], train_loss: 1761.9189, train_reconst: 1585.5430, train_u: 16.3333,train_z: 5.7242,train_c: 151.3896,train_l: 2.9288
Epoch[70/200], train_loss: 1755.7707, train_reconst: 1580.6499, train_u: 16.1848,train_z: 5.6212,train_c: 150.3357,train_l: 2.9791
Epoch[80/200], train_loss: 1748.3032, train_reconst: 1575.9084, train_u: 16.0641,train_z: 5.6020,train_c: 147.7620,train_l: 2.9667
Epoch[90/200], train_loss: 1745.4689, train_reconst: 1573.8276, train_u: 15.9672,train_z: 5.5890,train_c: 147.1002,train_l: 2.9849
Epoch[100/200], train_loss: 1741.4059, train_reconst: 1571.2467, train_u: 15.8885,train_z: 5.5138,train_c: 145.7818,train_l: 2.9751
Epoch[110/200], train_loss: 1739.2005, train_reconst: 1569.5755, train_u: 15.8249,train_z: 5.5090,train_c: 145.3119,train_l: 2.9792
Epoch[120/200], train_loss: 1737.3613, train_reconst: 1568.0982, train_u: 15.7730,train_z: 5.5225,train_c: 144.9962,train_l: 2.9714
Epoch[130/200], train_loss: 1735.5456, train_reconst: 1567.2170, train_u: 15.7308,train_z: 5.5234,train_c: 144.1060,train_l: 2.9684
Epoch[140/200], train_loss: 1733.8153, train_reconst: 1565.6691, train_u: 15.6964,train_z: 5.5186,train_c: 143.9690,train_l: 2.9622
Epoch[150/200], train_loss: 1734.0409, train_reconst: 1565.6646, train_u: 15.6686,train_z: 5.5101,train_c: 144.2291,train_l: 2.9684
Epoch[160/200], train_loss: 1733.5203, train_reconst: 1564.8707, train_u: 15.6460,train_z: 5.5183,train_c: 144.5110,train_l: 2.9744
Epoch[170/200], train_loss: 1731.6396, train_reconst: 1563.7964, train_u: 15.6274,train_z: 5.5189,train_c: 143.7256,train_l: 2.9713
Epoch[180/200], train_loss: 1730.6814, train_reconst: 1563.4632, train_u: 15.6123,train_z: 5.4934,train_c: 143.1347,train_l: 2.9779
Epoch[190/200], train_loss: 1730.8846, train_reconst: 1563.2024, train_u: 15.6000,train_z: 5.5424,train_c: 143.5656,train_l: 2.9743
Epoch[200/200], train_loss: 1730.3862, train_reconst: 1563.3851, train_u: 15.5899,train_z: 5.4530,train_c: 142.9817,train_l: 2.9766
Epoch[10/200], train_loss: 1964.5555, train_reconst: 1717.3358, train_u: 17.7731,train_z: 12.2491,train_c: 216.1686,train_l: 1.0289
Epoch[20/200], train_loss: 1836.5867, train_reconst: 1631.5397, train_u: 17.3562,train_z: 8.2599,train_c: 177.4633,train_l: 1.9675
Epoch[30/200], train_loss: 1803.6228, train_reconst: 1611.8570, train_u: 17.0200,train_z: 6.8935,train_c: 165.4288,train_l: 2.4234
Epoch[40/200], train_loss: 1784.1136, train_reconst: 1599.9807, train_u: 16.7433,train_z: 6.1974,train_c: 158.4979,train_l: 2.6943
Epoch[50/200], train_loss: 1771.6689, train_reconst: 1591.6377, train_u: 16.5159,train_z: 5.8556,train_c: 154.8143,train_l: 2.8453
Epoch[60/200], train_loss: 1761.9189, train_reconst: 1585.5430, train_u: 16.3333,train_z: 5.7242,train_c: 151.3896,train_l: 2.9288
Epoch[70/200], train_loss: 1755.7707, train_reconst: 1580.6499, train_u: 16.1848,train_z: 5.6212,train_c: 150.3357,train_l: 2.9791
Epoch[80/200], train_loss: 1748.3032, train_reconst: 1575.9084, train_u: 16.0641,train_z: 5.6020,train_c: 147.7620,train_l: 2.9667
Epoch[90/200], train_loss: 1745.4689, train_reconst: 1573.8276, train_u: 15.9672,train_z: 5.5890,train_c: 147.1002,train_l: 2.9849
Epoch[100/200], train_loss: 1741.4059, train_reconst: 1571.2467, train_u: 15.8885,train_z: 5.5138,train_c: 145.7818,train_l: 2.9751
Epoch[110/200], train_loss: 1739.2005, train_reconst: 1569.5755, train_u: 15.8249,train_z: 5.5090,train_c: 145.3119,train_l: 2.9792
Epoch[120/200], train_loss: 1737.3613, train_reconst: 1568.0982, train_u: 15.7730,train_z: 5.5225,train_c: 144.9962,train_l: 2.9714
Epoch[130/200], train_loss: 1735.5456, train_reconst: 1567.2170, train_u: 15.7308,train_z: 5.5234,train_c: 144.1060,train_l: 2.9684
Epoch[140/200], train_loss: 1733.8153, train_reconst: 1565.6691, train_u: 15.6964,train_z: 5.5186,train_c: 143.9690,train_l: 2.9622
Epoch[150/200], train_loss: 1734.0409, train_reconst: 1565.6646, train_u: 15.6686,train_z: 5.5101,train_c: 144.2291,train_l: 2.9684
Epoch[160/200], train_loss: 1733.5203, train_reconst: 1564.8707, train_u: 15.6460,train_z: 5.5183,train_c: 144.5110,train_l: 2.9744
Epoch[170/200], train_loss: 1731.6396, train_reconst: 1563.7964, train_u: 15.6274,train_z: 5.5189,train_c: 143.7256,train_l: 2.9713
Epoch[180/200], train_loss: 1730.6814, train_reconst: 1563.4632, train_u: 15.6123,train_z: 5.4934,train_c: 143.1347,train_l: 2.9779
Epoch[190/200], train_loss: 1730.8846, train_reconst: 1563.2024, train_u: 15.6000,train_z: 5.5424,train_c: 143.5656,train_l: 2.9743
Epoch[200/200], train_loss: 1730.3862, train_reconst: 1563.3851, train_u: 15.5899,train_z: 5.4530,train_c: 142.9817,train_l: 2.9766
Epoch[10/200], train_loss: 1964.5555, train_reconst: 1717.3358, train_u: 17.7731,train_z: 12.2491,train_c: 216.1686,train_l: 1.0289
Epoch[20/200], train_loss: 1836.5867, train_reconst: 1631.5397, train_u: 17.3562,train_z: 8.2599,train_c: 177.4633,train_l: 1.9675
Epoch[30/200], train_loss: 1803.6228, train_reconst: 1611.8570, train_u: 17.0200,train_z: 6.8935,train_c: 165.4288,train_l: 2.4234
Epoch[40/200], train_loss: 1784.1136, train_reconst: 1599.9807, train_u: 16.7433,train_z: 6.1974,train_c: 158.4979,train_l: 2.6943
Epoch[50/200], train_loss: 1771.6689, train_reconst: 1591.6377, train_u: 16.5159,train_z: 5.8556,train_c: 154.8143,train_l: 2.8453
Epoch[60/200], train_loss: 1761.9189, train_reconst: 1585.5430, train_u: 16.3333,train_z: 5.7242,train_c: 151.3896,train_l: 2.9288
Epoch[70/200], train_loss: 1755.7707, train_reconst: 1580.6499, train_u: 16.1848,train_z: 5.6212,train_c: 150.3357,train_l: 2.9791
Epoch[80/200], train_loss: 1748.3032, train_reconst: 1575.9084, train_u: 16.0641,train_z: 5.6020,train_c: 147.7620,train_l: 2.9667
Epoch[90/200], train_loss: 1745.4689, train_reconst: 1573.8276, train_u: 15.9672,train_z: 5.5890,train_c: 147.1002,train_l: 2.9849
Epoch[100/200], train_loss: 1741.4059, train_reconst: 1571.2467, train_u: 15.8885,train_z: 5.5138,train_c: 145.7818,train_l: 2.9751
Epoch[110/200], train_loss: 1739.2005, train_reconst: 1569.5755, train_u: 15.8249,train_z: 5.5090,train_c: 145.3119,train_l: 2.9792
Epoch[120/200], train_loss: 1737.3613, train_reconst: 1568.0982, train_u: 15.7730,train_z: 5.5225,train_c: 144.9962,train_l: 2.9714
Epoch[130/200], train_loss: 1735.5456, train_reconst: 1567.2170, train_u: 15.7308,train_z: 5.5234,train_c: 144.1060,train_l: 2.9684
Epoch[140/200], train_loss: 1733.8153, train_reconst: 1565.6691, train_u: 15.6964,train_z: 5.5186,train_c: 143.9690,train_l: 2.9622
Epoch[150/200], train_loss: 1734.0409, train_reconst: 1565.6646, train_u: 15.6686,train_z: 5.5101,train_c: 144.2291,train_l: 2.9684
Epoch[160/200], train_loss: 1733.5203, train_reconst: 1564.8707, train_u: 15.6460,train_z: 5.5183,train_c: 144.5110,train_l: 2.9744
Epoch[170/200], train_loss: 1731.6396, train_reconst: 1563.7964, train_u: 15.6274,train_z: 5.5189,train_c: 143.7256,train_l: 2.9713
Epoch[180/200], train_loss: 1730.6814, train_reconst: 1563.4632, train_u: 15.6123,train_z: 5.4934,train_c: 143.1347,train_l: 2.9779
Epoch[190/200], train_loss: 1730.8846, train_reconst: 1563.2024, train_u: 15.6000,train_z: 5.5424,train_c: 143.5656,train_l: 2.9743
Epoch[200/200], train_loss: 1730.3862, train_reconst: 1563.3851, train_u: 15.5899,train_z: 5.4530,train_c: 142.9817,train_l: 2.9766
✓ starfysh is done
The starfysh result is saved in self.adata_cell2location
The starfysh model is saved in self.starfysh_model
Step 4: Visualization (5–15 min)¶
We provide multiple views: single-target spatial heatmaps, multi-target overlays, and local pie charts. Start with global inspection, then zoom into biologically relevant regions for higher-resolution assessment.
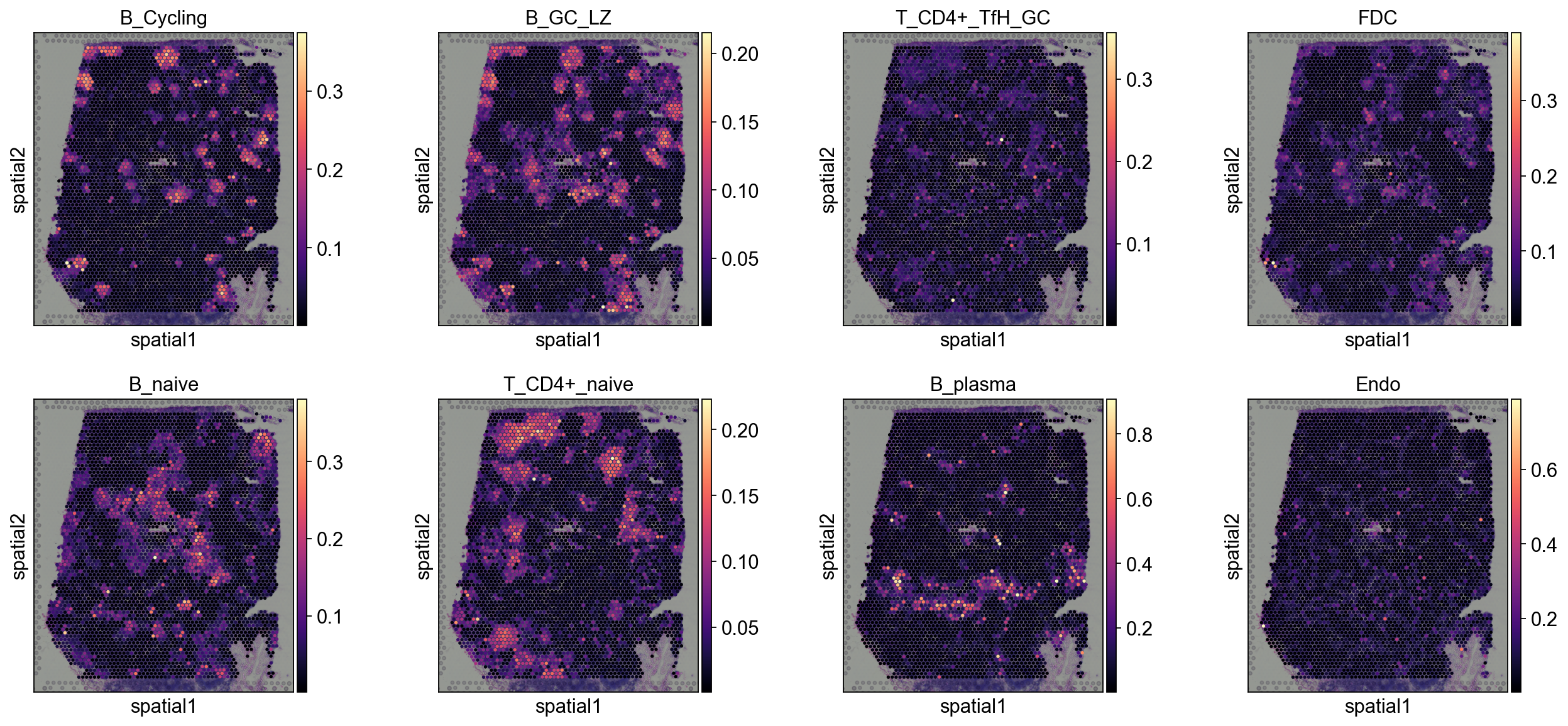
4.1 Spatial value dotplot¶
annotation_list=['B_Cycling', 'B_GC_LZ', 'T_CD4+_TfH_GC', 'FDC',
'B_naive', 'T_CD4+_naive', 'B_plasma', 'Endo']
sc.pl.spatial(
decov_obj.adata_cell2location,
cmap='magma',
# show first 8 cell types
color=annotation_list,
ncols=4, size=1.3,
img_key='hires',
# limit color scale at 99.2% quantile of cell abundance
#vmin=0, vmax='p99.2'
)
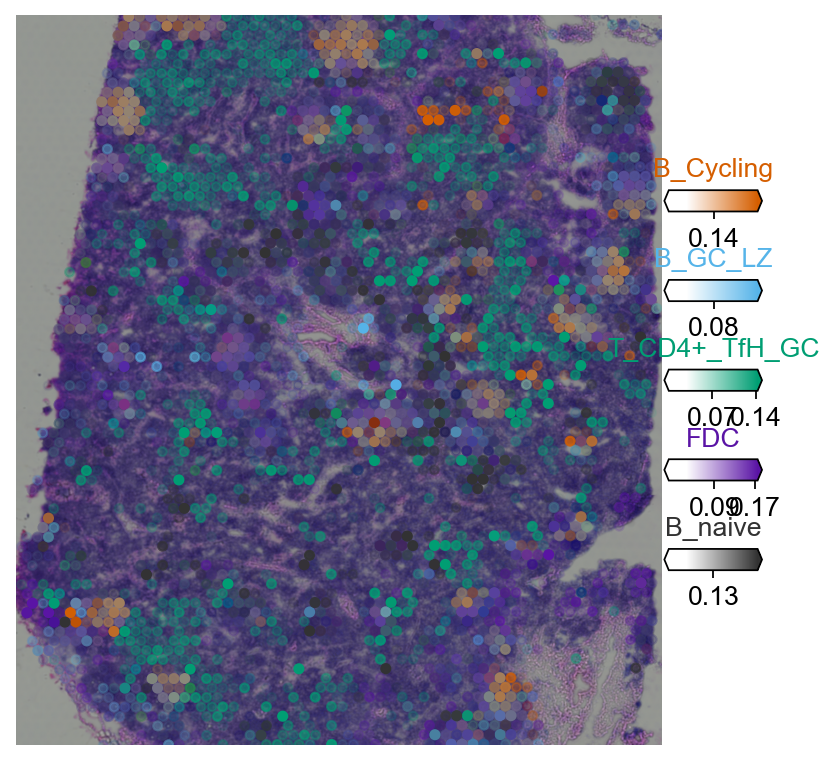
4.2 Multi-target overlay¶
import matplotlib as mpl
clust_labels = annotation_list[:5]
clust_col = ['' + str(i) for i in clust_labels] # in case column names differ from labels
with mpl.rc_context({'figure.figsize': (6, 6),'axes.grid': False}):
fig = ov.pl.plot_spatial(
adata=decov_obj.adata_cell2location,
# labels to show on a plot
color=clust_col, labels=clust_labels,
show_img=True,
# 'fast' (white background) or 'dark_background'
style='fast',
# limit color scale at 99.2% quantile of cell abundance
max_color_quantile=0.992,
# size of locations (adjust depending on figure size)
circle_diameter=4,
reorder_cmap = [#0,
1,2,3,4,6], #['yellow', 'orange', 'blue', 'green', 'purple', 'grey', 'white'],
colorbar_position='right',
#palette=color_dict
)
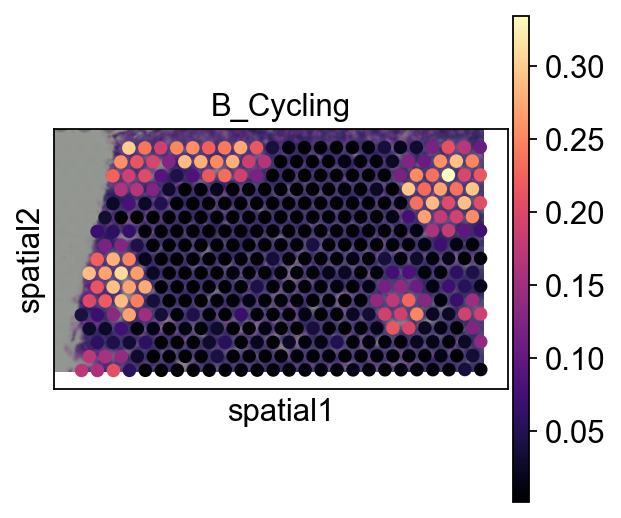
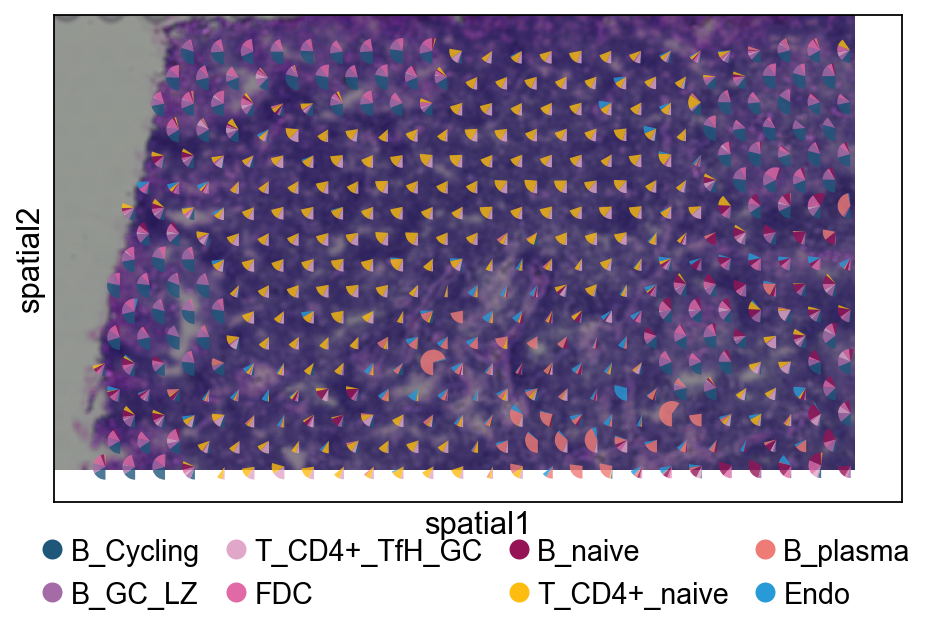
4.3 Pie plot¶
We recommend cropping a region of interest before plotting to avoid overly dense pie charts on whole slides.
adata_s = ov.space.crop_space_visium(
decov_obj.adata_cell2location,
crop_loc=(0, 0),
crop_area=(500, 1000),
library_id=list(decov_obj.adata_cell2location.uns['spatial'].keys())[0] ,
scale=1
)
sc.pl.spatial(adata_s, cmap='magma',
# show first 8 cell types
color=annotation_list[0],
ncols=4, size=1.3,
img_key='hires',
# limit color scale at 99.2% quantile of cell abundance
#vmin=0, vmax='p99.2'
)
color_dict=dict(zip(annotation_list,
ov.pl.sc_color))
fig, ax = ov.plt.subplots(figsize=(8, 4))
sc.pl.spatial(
adata_s,
basis='spatial',
color=None,
size=1.3,
img_key='hires',
ax=ax,
show=False
)
ov.pl.add_pie2spatial(
adata_s,
img_key='hires',
cell_type_columns=annotation_list[:],
ax=ax,
colors=color_dict,
pie_radius=10,
remainder='gap',
legend_loc=(0.5, -0.25),
ncols=4,
alpha=0.8
)
#plt.show()