Automatic cell type annotation with GPT/Other¶
GPTCelltype, an open-source R software package to facilitate cell type annotation by GPT-4.
We made three improvements in integrating the GPTCelltype algorithm in OmicVerse:
Native support for Python: Since GPTCelltype is an R language package, in order to make it conform to scverse’s anndata ecosystem, we have rewritten the whole function so that it works perfectly under Python.
More model support: We provide more big models to choose from outside of Openai, e.g. Qwen(通义千问), Kimi, and more model support is available through the parameter
base_urlormodel_name.
If you found this tutorial helpful, please cite GPTCelltype and OmicVerse:
Hou, W. and Ji, Z., 2023. Reference-free and cost-effective automated cell type annotation with GPT-4 in single-cell RNA-seq analysis. Nature Methods, 2024 March 25.
import omicverse as ov
print(f'omicverse version:{ov.__version__}')
import scanpy as sc
print(f'scanpy version:{sc.__version__}')
ov.ov_plot_set()
____ _ _ __
/ __ \____ ___ (_)___| | / /__ _____________
/ / / / __ `__ \/ / ___/ | / / _ \/ ___/ ___/ _ \
/ /_/ / / / / / / / /__ | |/ / __/ / (__ ) __/
\____/_/ /_/ /_/_/\___/ |___/\___/_/ /____/\___/
Version: 1.6.2, Tutorials: https://omicverse.readthedocs.io/
omicverse version:1.6.2
scanpy version:1.10.1
Loading data¶
The data consist of 3k PBMCs from a Healthy Donor and are freely available from 10x Genomics (here from this webpage). On a unix system, you can uncomment and run the following to download and unpack the data. The last line creates a directory for writing processed data.
# !mkdir data
# !wget http://cf.10xgenomics.com/samples/cell-exp/1.1.0/pbmc3k/pbmc3k_filtered_gene_bc_matrices.tar.gz -O data/pbmc3k_filtered_gene_bc_matrices.tar.gz
# !cd data; tar -xzf pbmc3k_filtered_gene_bc_matrices.tar.gz
# !mkdir write
Read in the count matrix into an AnnData object, which holds many slots for annotations and different representations of the data. It also comes with its own HDF5-based file format: .h5ad.
adata = sc.read_10x_mtx(
'data/filtered_gene_bc_matrices/hg19/', # the directory with the `.mtx` file
var_names='gene_symbols', # use gene symbols for the variable names (variables-axis index)
cache=True) # write a cache file for faster subsequent reading
... reading from cache file cache/data-filtered_gene_bc_matrices-hg19-matrix.h5ad
Data preprocessing¶
Here, we use ov.single.scanpy_lazy to preprocess the raw data of scRNA-seq, it included filter the doublets cells, normalizing counts per cell, log1p, extracting highly variable genes, and cluster of cells calculation.
But if you want to experience step-by-step preprocessing, we also provide more detailed preprocessing steps here, please refer to our preprocess chapter for a detailed explanation.
We stored the raw counts in count layers, and the raw data in adata.raw.to_adata().
#adata=ov.single.scanpy_lazy(adata)
#quantity control
adata=ov.pp.qc(adata,
tresh={'mito_perc': 0.05, 'nUMIs': 500, 'detected_genes': 250})
#normalize and high variable genes (HVGs) calculated
adata=ov.pp.preprocess(adata,mode='shiftlog|pearson',n_HVGs=2000,)
#save the whole genes and filter the non-HVGs
adata.raw = adata
adata = adata[:, adata.var.highly_variable_features]
#scale the adata.X
ov.pp.scale(adata)
#Dimensionality Reduction
ov.pp.pca(adata,layer='scaled',n_pcs=50)
#Neighbourhood graph construction
sc.pp.neighbors(adata, n_neighbors=15, n_pcs=50,
use_rep='scaled|original|X_pca')
#clusters
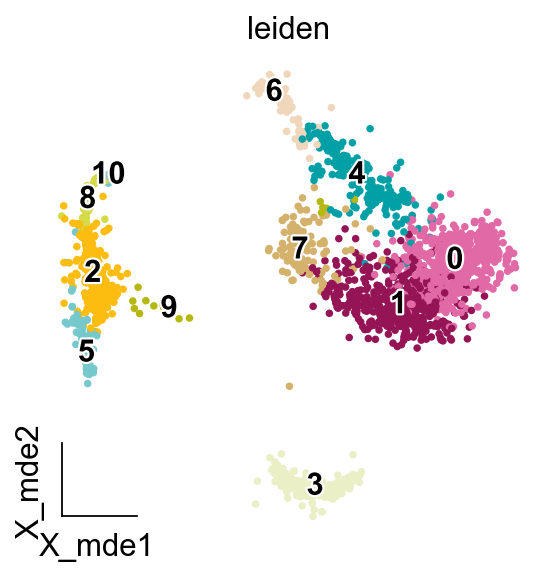
sc.tl.leiden(adata)
#find marker
sc.tl.dendrogram(adata,'leiden',use_rep='scaled|original|X_pca')
sc.tl.rank_genes_groups(adata, 'leiden', use_rep='scaled|original|X_pca',
method='wilcoxon',use_raw=False,)
#Dimensionality Reduction for visualization(X_mde=X_umap+GPU)
adata.obsm["X_mde"] = ov.utils.mde(adata.obsm["scaled|original|X_pca"])
adata
GPT Celltype¶
gptcelltype supports dictionary format input, we provide omicverse.single.get_celltype_marker to get the marker genes for each cell type as a dictionary.
Using genes manually¶
We can manually define a dictionary to determine the accuracy of the output
import os
all_markers={'cluster1':['CD3D','CD3E'],
'cluster2':['MS4A1']}
os.environ['AGI_API_KEY'] = 'sk-**' # Replace with your actual API key
result = ov.single.gptcelltype(all_markers, tissuename='PBMC', speciename='human',
model='qwen-plus', provider='qwen',
topgenenumber=5)
result
Note: AGI API key found: returning the cell type annotations.
Note: It is always recommended to check the results returned by GPT-4 in case of AI hallucination, before going to downstream analysis.
{'cluster1': '1. T cells (CD3D, CD3E)', 'cluster2': '2. B cells (MS4A1)'}
Get Genes for Each Cluster Automatically¶
all_markers=ov.single.get_celltype_marker(adata,clustertype='leiden',rank=True,
key='rank_genes_groups',
foldchange=2,topgenenumber=5)
all_markers
...get cell type marker
{'0': ['LTB', 'LDHB', 'IL32', 'CD3D', 'IL7R'],
'1': ['LDHB', 'CD3D', 'NOSIP', 'CD3E', 'C6orf48'],
'10': ['SPARC', 'PPBP', 'GNG11', 'PF4', 'CD9'],
'2': ['LYZ', 'S100A9', 'S100A8', 'FCN1', 'TYROBP'],
'3': ['CD74', 'CD79A', 'HLA-DRA', 'CD79B', 'HLA-DPB1'],
'4': ['CCL5', 'NKG7', 'CST7', 'GZMA', 'IL32'],
'5': ['LST1', 'FCER1G', 'COTL1', 'AIF1', 'IFITM3'],
'6': ['NKG7', 'GZMB', 'GNLY', 'CTSW', 'PRF1'],
'7': ['CCL5', 'GZMK'],
'8': ['FTL', 'FTH1', 'S100A8', 'S100A9', 'TYROBP'],
'9': ['HLA-DRB1', 'HLA-DPA1', 'HLA-DPB1', 'HLA-DRA', 'HLA-DRB5']}
Option 1. Through OpenAI API (provider or base_url)¶
Use ov.single.gptcelltype function to annotate cell types.
You can simply set provider (or base_url) and provider parameters to provide the function with base url and the exact model which are required for model calling.
import os
os.environ['AGI_API_KEY'] = 'sk-**' # Replace with your actual API key
result = ov.single.gptcelltype(all_markers, tissuename='PBMC', speciename='human',
model='qwen-plus', provider='qwen',
topgenenumber=5)
result
Note: AGI API key found: returning the cell type annotations.
Note: It is always recommended to check the results returned by GPT-4 in case of AI hallucination, before going to downstream analysis.
{'0': '1. T cells (CD3D, IL7R)',
'1': '2. T cells (CD3D, CD3E)',
'10': '3. Platelets or Megakaryocytes (CD9)',
'2': '4. Neutrophils (LYZ, S100A9, S100A8, FCN1, TYROBP)',
'3': '5. B cells (CD74, CD79A, HLA-DRA, CD79B, HLA-DPB1)',
'4': '6. Cytotoxic T cells or NK cells (NKG7, CST7, GZMA, IL32)',
'5': '7. Monocytes or Macrophages (LST1, FCER1G, COTL1, AIF1, IFITM3)',
'6': '8. Cytotoxic T cells or NK cells (NKG7, GZMB, GNLY, CTSW, PRF1)',
'7': '9. Cytotoxic T cells (GZMK)',
'8': '10. Neutrophils (FTL, FTH1, S100A8, S100A9, TYROBP)',
'9': '11. Antigen presentation (HLA-DRB1, HLA-DPA1, HLA-DPB1, HLA-DRA, HLA-DRB5)'}
We can keep only the cell type of the output and remove other irrelevant information.
new_result={}
for key in result.keys():
new_result[key]=result[key].split(': ')[-1].split(' (')[0].split('. ')[1]
new_result
{'0': 'T cells',
'1': 'T cells',
'10': 'Platelets or Megakaryocytes',
'2': 'Neutrophils',
'3': 'B cells',
'4': 'Cytotoxic T cells or NK cells',
'5': 'Monocytes or Macrophages',
'6': 'Cytotoxic T cells or NK cells',
'7': 'Cytotoxic T cells',
'8': 'Neutrophils',
'9': 'Antigen presentation'}
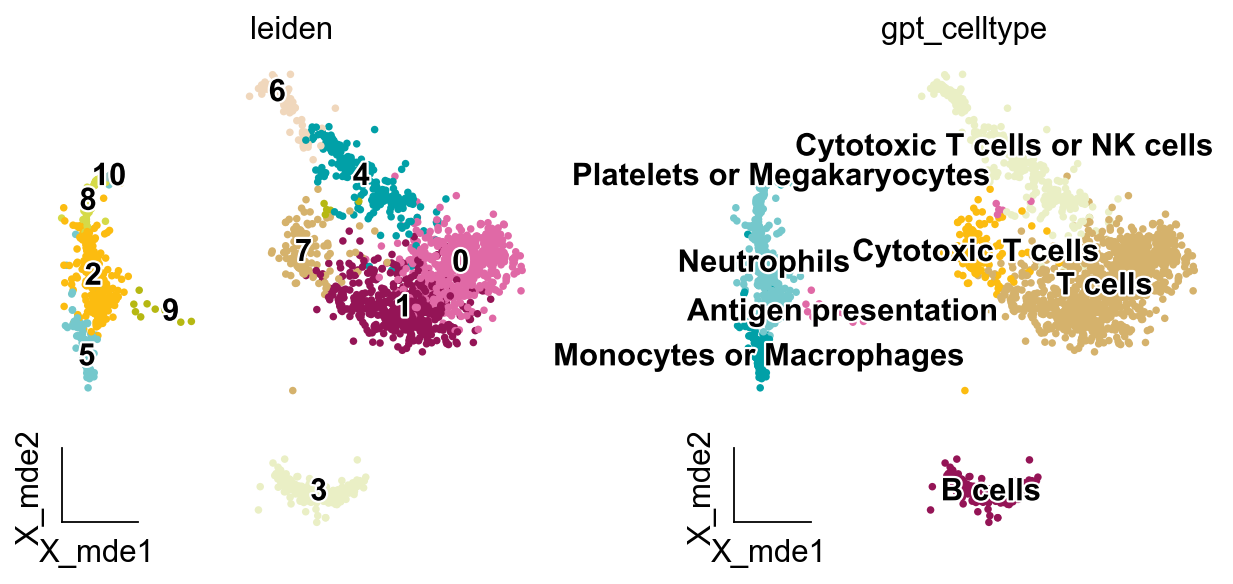
adata.obs['gpt_celltype'] = adata.obs['leiden'].map(new_result).astype('category')
More models¶
Our implementation of gptcelltype in omicverse supports almost all large language models that support the openai api format.
all_markers={'cluster1':['CD3D','CD3E'],
'cluster2':['MS4A1']}
Openai¶
The OpenAI API uses API keys for authentication. You can create API keys at a user or service account level. Service accounts are tied to a “bot” individual and should be used to provision access for production systems. Each API key can be scoped to one of the following,
User keys - Our legacy keys. Provides access to all organizations and all projects that user has been added to; access API Keys to view your available keys. We highly advise transitioning to project keys for best security practices, although access via this method is currently still supported.
Please select the model you need to use: list of supported models.
os.environ['AGI_API_KEY'] = 'sk-**' # Replace with your actual API key
result = ov.single.gptcelltype(all_markers, tissuename='PBMC', speciename='human',
model='gpt-4o', provider='openai',
topgenenumber=5)
result
Note: AGI API key found: returning the cell type annotations.
Note: It is always recommended to check the results returned by GPT-4 in case of AI hallucination, before going to downstream analysis.
{'cluster1': '1. T cell', 'cluster2': '2. B cell'}
Qwen(通义千问)¶
Enabled DashScope service and obtained API-KEY: Enabled DashScope and created API-KEY.
We recommend you to configure API-KEY in environment variable to reduce the risk of API-KEY leakage, please refer to Configuring API-KEY through Environment Variable for the configuration method, you can also configure API-KEY in code, but the risk of leakage will be increased.
Please select the model you need to use: list of supported models.
简体中文
已开通灵积模型服务并获得API-KEY:开通DashScope并创建API-KEY。
我们推荐您将API-KEY配置到环境变量中以降低API-KEY的泄漏风险,配置方法可参考通过环境变量配置API-KEY。您也可以在代码中配置API-KEY,但是泄漏风险会提高。
请选择您需要使用的模型:支持的模型列表。
os.environ['AGI_API_KEY'] = 'sk-**' # Replace with your actual API key
result = ov.single.gptcelltype(all_markers, tissuename='PBMC', speciename='human',
model='qwen-plus', provider='qwen',
topgenenumber=5)
result
Note: AGI API key found: returning the cell type annotations.
Note: It is always recommended to check the results returned by GPT-4 in case of AI hallucination, before going to downstream analysis.
{'cluster1': 'CD3D, CD3E: T cells', 'cluster2': 'MS4A1: B cells'}
Kimi(月之暗面)¶
You will need a Dark Side of the Moon API key to use our service. You can create an API key in Console.
Please select the model you need to use: List of supported models
简体中文
os.environ['AGI_API_KEY'] = 'sk-**' # Replace with your actual API key
result = ov.single.gptcelltype(all_markers, tissuename='PBMC', speciename='human',
model='moonshot-v1-8k', provider='kimi',
topgenenumber=5)
result
Note: AGI API key found: returning the cell type annotations.
Note: It is always recommended to check the results returned by GPT-4 in case of AI hallucination, before going to downstream analysis.
{'cluster1': '1. T cell', 'cluster2': '2. B cell'}
Other Models¶
You can manually set the base_url parameter to specify other models that need to be used, note that the model needs to support Openai’s parameters. Three examples are provided here (when you specify the base_url parameter, the provider parameter will be invalid):
if provider == 'openai':
base_url = "https://api.openai.com/v1/"
elif provider == 'kimi':
base_url = "https://api.moonshot.cn/v1"
elif provider == 'qwen':
base_url = "https://dashscope.aliyuncs.com/compatible-mode/v1"
os.environ['AGI_API_KEY'] = 'sk-**' # Replace with your actual API key
result = ov.single.gptcelltype(all_markers, tissuename='PBMC', speciename='human',
model='moonshot-v1-8k', base_url="https://api.moonshot.cn/v1",
topgenenumber=5)
result
Note: AGI API key found: returning the cell type annotations.
Note: It is always recommended to check the results returned by GPT-4 in case of AI hallucination, before going to downstream analysis.
{'cluster1': '1. T cell', 'cluster2': '2. B cell'}
Option 2. Through Local LLM (model_name)¶
Use ov.single.gptcelltype_local function to annotate cell types.
You can simply set the model_name parameter.
anno_model = 'path/to/your/local/LLM' # '~/models/Qwen2-7B-Instruct'
result = ov.single.gptcelltype_local(all_markers, tissuename='PBMC', speciename='human',
model_name=anno_model, topgenenumber=5)
result
0: B cells [CD3D, IL7R]
1: Cytotoxic T cells [CCL5, NKG7, GZMA]; Natural Killer (NK) cells [CD3D, CD3E]
10: Endothelial cells [SPARC, PPBP]; Platelets [PF4]
2: Neutrophils [LYZ, S100A9, S100A8]; Monocytes [FCN1]
3: B cells [CD74, CD79A, HLA-DRA, CD79B, HLA-DPB1]
4: Cytotoxic T cells [CCL5, NKG7, CST7, GZMA, IL32]
5: DNA repair cells [LST1, FCER1G, COTL1, AIF1]; Cytotoxic T cells [IFITM3]
6: Cytotoxic T cells [NKG7, GZMB, GNLY, CTSW, PRF1]
7: Cytotoxic T cells [CCL5, GZMK]
8: Erythroid progenitors [FTL, FTH1, S100A8, S100A9]; Cytotoxic T cells [TYROBP]
9: Dendritic cells [HLA-DRB1, HLA-DPA1, HLA-DPB1, HLA-DRA, HLA-DRB5]
Note: It is always recommended to check the results returned by the LLM in case of AI hallucination, before going to downstream analysis.
{'0': 'B cells',
'1': 'Cytotoxic T cells',
'10': 'Endothelial cells',
'2': 'Neutrophils',
'3': 'B cells',
'4': 'Cytotoxic T cells',
'5': 'DNA repair cells',
'6': 'Cytotoxic T cells',
'7': 'Cytotoxic T cells',
'8': 'Erythroid progenitors',
'9': 'Dendritic cells'}
Note that you may encounter network problems that prevent you from downloading LLMs.
In this case, please refer to https://zhuanlan.zhihu.com/p/663712983