Alignment and RNA velocity analysis of single-cell RNA-seq data.¶
The kallisto, bustools and kb-python programs are free, open-source software tools for performing this analysis that together can produce gene expression quantification from raw sequencing reads. In this tutorial, we will perform pre-processing and RNA velocity analysis of human week 10 fetal forebrain dataset (SRR6470906) from La Manno et al., 2018 using the kallisto | bustools workflow.
We made an improvement in integrating the kallisto, bustools and kb-python program in OmicVerse:
More user-friendly function implementation: we automated their encapsulation into the
omicverse.alignmentclass.
If you found this tutorial helpful, please cite kb-python and OmicVerse:
Sullivan, D.K., Min, K.H.(., Hjörleifsson, K.E. et al. kallisto, bustools and kb-python for quantifying bulk, single-cell and single-nucleus RNA-seq. Nature Protocol (2025). https://doi.org/10.1038/s41596-024-01057-0
Melsted, P., Booeshaghi, A.S., Liu, L. et al. Modular, efficient and constant-memory single-cell RNA-seq preprocessing. Nature Biotechnology (2021). https://doi.org/10.1038/s41587-021-00870-2
import omicverse as ov
import scanpy as sc
import pandas as pd
import numpy as np
ov.plot_set(font_path='Arial')
🔬 Starting plot initialization...
Using already downloaded Arial font from: /tmp/omicverse_arial.ttf
Registered as: Arial
🧬 Detecting GPU devices…
✅ NVIDIA CUDA GPUs detected: 8
• [CUDA 0] NVIDIA GeForce RTX 4090 D
Memory: 23.6 GB | Compute: 8.9
• [CUDA 1] NVIDIA GeForce RTX 4090 D
Memory: 23.6 GB | Compute: 8.9
• [CUDA 2] NVIDIA GeForce RTX 4090 D
Memory: 23.6 GB | Compute: 8.9
• [CUDA 3] NVIDIA GeForce RTX 4090 D
Memory: 23.6 GB | Compute: 8.9
• [CUDA 4] NVIDIA GeForce RTX 4090 D
Memory: 23.6 GB | Compute: 8.9
• [CUDA 5] NVIDIA GeForce RTX 4090 D
Memory: 23.6 GB | Compute: 8.9
• [CUDA 6] NVIDIA GeForce RTX 4090 D
Memory: 23.6 GB | Compute: 8.9
• [CUDA 7] NVIDIA GeForce RTX 4090 D
Memory: 23.6 GB | Compute: 8.9
✅ plot_set complete.
Download human reference files and build the index¶
We build a human cDNA and intron index from the human genome and annotations provided by Ensembl.
%%time
!wget -P velocity ftp://ftp.ensembl.org/pub/release-108/fasta/homo_sapiens/dna/Homo_sapiens.GRCh38.dna.primary_assembly.fa.gz
!wget -P velocity ftp://ftp.ensembl.org/pub/release-108/gtf/homo_sapiens/Homo_sapiens.GRCh38.108.gtf.gz
--2025-10-24 06:42:54-- ftp://ftp.ensembl.org/pub/release-108/fasta/homo_sapiens/dna/Homo_sapiens.GRCh38.dna.primary_assembly.fa.gz
=> ‘velocity/Homo_sapiens.GRCh38.dna.primary_assembly.fa.gz.1’
Resolving ftp.ensembl.org (ftp.ensembl.org)... ^C
--2025-10-24 06:42:57-- ftp://ftp.ensembl.org/pub/release-108/gtf/homo_sapiens/Homo_sapiens.GRCh38.108.gtf.gz
=> ‘velocity/Homo_sapiens.GRCh38.108.gtf.gz.1’
Resolving ftp.ensembl.org (ftp.ensembl.org)... 193.62.193.169
Connecting to ftp.ensembl.org (ftp.ensembl.org)|193.62.193.169|:21... connected.
Logging in as anonymous ... Logged in!
==> SYST ... done. ==> PWD ... done.
==> TYPE I ... done. ==> CWD (1) /pub/release-108/gtf/homo_sapiens ... done.
==> SIZE Homo_sapiens.GRCh38.108.gtf.gz ... 54107597
==> PASV ... done. ==> RETR Homo_sapiens.GRCh38.108.gtf.gz ... ^C
CPU times: user 2.13 s, sys: 404 ms, total: 2.54 s
Wall time: 5.89 s
Here we build a human RNA velocity index, and you can see this notebook for more details about how to RNA velocity indices.
result=ov.alignment.single.ref(
fasta_paths='velocity/Homo_sapiens.GRCh38.dna.primary_assembly.fa.gz', #input
gtf_paths='velocity/Homo_sapiens.GRCh38.108.gtf.gz', #input
index_path='velocity/index.idx', #output
t2g_path='velocity/t2g.txt', #output
cdna_path='velocity/cdna.fa', #output
f2='velocity/intron.fa', #output
c1='velocity/cdna_t2c.txt', #output
c2='velocity/intron_t2c.txt', #output
workflow='lamanno',
threads=8
)
print(result.keys())
🚀 Starting ref workflow: lamanno
Using temporary directory: tmp-kb-06ff3e80a53545f3920a4681bc45da78
>> /opt/miniforge/envs/omicverse_working/bin/kb ref --workflow lamanno --tmp tmp-kb-06ff3e80a53545f3920a4681bc45da78 -i velocity/index.idx -g velocity/t2g.txt -t 8 --d-list-overhang 1 -f1 velocity/cdna.fa -f2 velocity/intron.fa -c1 velocity/cdna_t2c.txt -c2 velocity/intron_t2c.txt velocity/Homo_sapiens.GRCh38.dna.primary_assembly.fa.gz velocity/Homo_sapiens.GRCh38.108.gtf.gz
[2025-10-29 07:41:52,381] INFO [ref_lamanno] Preparing velocity/Homo_sapiens.GRCh38.dna.primary_assembly.fa.gz, velocity/Homo_sapiens.GRCh38.108.gtf.gz
[2025-10-29 07:42:56,969] INFO [ref_lamanno] Splitting genome velocity/Homo_sapiens.GRCh38.dna.primary_assembly.fa.gz into cDNA at /data/hulei/Projects/Omicverse_2/omicverse_update/tmp-kb-06ff3e80a53545f3920a4681bc45da78/tmpd6_5n02u
[2025-10-29 07:44:26,416] INFO [ref_lamanno] Creating cDNA transcripts-to-capture at /data/hulei/Projects/Omicverse_2/omicverse_update/tmp-kb-06ff3e80a53545f3920a4681bc45da78/tmp4uoqoscv
[2025-10-29 07:44:30,592] INFO [ref_lamanno] Splitting genome into introns at /data/hulei/Projects/Omicverse_2/omicverse_update/tmp-kb-06ff3e80a53545f3920a4681bc45da78/tmpuqw6zk3j
[2025-10-29 07:53:31,728] INFO [ref_lamanno] Creating intron transcripts-to-capture at /data/hulei/Projects/Omicverse_2/omicverse_update/tmp-kb-06ff3e80a53545f3920a4681bc45da78/tmppqqiuleo
[2025-10-29 07:55:05,492] INFO [ref_lamanno] Concatenating 1 cDNA FASTAs to velocity/cdna.fa
[2025-10-29 07:55:09,790] INFO [ref_lamanno] Concatenating 1 cDNA transcripts-to-captures to velocity/cdna_t2c.txt
[2025-10-29 07:55:09,870] INFO [ref_lamanno] Concatenating 1 intron FASTAs to velocity/intron.fa
[2025-10-29 07:56:41,579] INFO [ref_lamanno] Concatenating 1 intron transcripts-to-captures to velocity/intron_t2c.txt
[2025-10-29 07:56:42,243] INFO [ref_lamanno] Concatenating cDNA and intron FASTAs to /data/hulei/Projects/Omicverse_2/omicverse_update/tmp-kb-06ff3e80a53545f3920a4681bc45da78/tmp11ndpu2o
[2025-10-29 07:59:43,573] INFO [ref_lamanno] Creating transcript-to-gene mapping at velocity/t2g.txt
[2025-10-29 08:01:22,179] INFO [ref_lamanno] Indexing /data/hulei/Projects/Omicverse_2/omicverse_update/tmp-kb-06ff3e80a53545f3920a4681bc45da78/tmp11ndpu2o to velocity/index.idx
✓ ref workflow completed!
dict_keys(['workflow', 'technology', 'parameters', 'index_path', 't2g_path', 'cdna_path'])
Generate RNA velocity count matrices¶
The following command will generate an RNA count matrix of cells (rows) by genes (columns) in H5AD format, which is a binary format used to store Anndata objects. Notice we are providing the index and transcript-to-gene mapping we downloaded in the previous step to the -i and -g arguments respectively, as well as the transcripts-to-capture lists to the -c1 and -c2 arguments. Also, these reads were generated with the 10x Genomics Chromium Single Cell v2 Chemistry, hence the -x 10xv2 argument.
!wget -P velocity https://caltech.box.com/shared/static/nvzqphhklk1yx938l6omursw7sr68y43.gz -O SRR6470906_S1_L001_R1_001.fastq.gz
!wget -P velocity https://caltech.box.com/shared/static/63fh2xa5t82x7s74rqa0e2u2ur59y5ox.gz -O SRR6470906_S1_L001_R2_001.fastq.gz
!wget -P velocity https://caltech.box.com/shared/static/zqi3durukillaw1pbns1kd1lowyfg5qk.gz -O SRR6470906_S1_L002_R1_001.fastq.gz
!wget -P velocity https://caltech.box.com/shared/static/i56qojfz41ns1kw9z86sla0vawsch96t.gz -O SRR6470906_S1_L002_R2_001.fastq.gz
result=ov.alignment.single.count(
fastq_paths=['velocity/SRR6470906_S1_L001_R1_001.fastq.gz', #input
'velocity/SRR6470906_S1_L001_R2_001.fastq.gz', #input
'velocity/SRR6470906_S1_L002_R1_001.fastq.gz', #input
'velocity/SRR6470906_S1_L002_R2_001.fastq.gz'],#input
output_path='velocity/SRR6470906', #input
index_path='velocity/index.idx', #input
t2g_path='velocity/t2g.txt', #input
c1='velocity/cdna_t2c.txt', #input
c2='velocity/intron_t2c.txt', #input
technology='10xv2',
workflow='lamanno',
h5ad=True,
filter_barcodes=True, #
)
print(result.keys())
🚀 Starting count workflow: lamanno
Technology: 10xv2
Output directory: velocity/SRR6470906
Using temporary directory: tmp-kb-8f35dcc278fb4900b238db3d4c120f14
>> /opt/miniforge/envs/omicverse_working/bin/kb count --workflow lamanno --tmp tmp-kb-8f35dcc278fb4900b238db3d4c120f14 -i velocity/index.idx -g velocity/t2g.txt -x 10xv2 -o velocity/SRR6470906 -c1 velocity/cdna_t2c.txt -c2 velocity/intron_t2c.txt -t 8 -m 2G --filter bustools --h5ad -c1 velocity/cdna_t2c.txt -c2 velocity/intron_t2c.txt velocity/SRR6470906_S1_L001_R1_001.fastq.gz velocity/SRR6470906_S1_L001_R2_001.fastq.gz velocity/SRR6470906_S1_L002_R1_001.fastq.gz velocity/SRR6470906_S1_L002_R2_001.fastq.gz
[2025-10-29 10:56:35,097] INFO [count_lamanno] Skipping kallisto bus because output files already exist. Use the --overwrite flag to overwrite.
[2025-10-29 10:56:35,097] INFO [count_lamanno] Sorting BUS file velocity/SRR6470906/output.bus to tmp-kb-8f35dcc278fb4900b238db3d4c120f14/output.s.bus
[2025-10-29 10:57:49,007] INFO [count_lamanno] On-list not provided
[2025-10-29 10:57:49,007] INFO [count_lamanno] Copying pre-packaged 10XV2 on-list to velocity/SRR6470906
[2025-10-29 10:57:49,188] INFO [count_lamanno] Inspecting BUS file tmp-kb-8f35dcc278fb4900b238db3d4c120f14/output.s.bus
[2025-10-29 10:57:53,696] INFO [count_lamanno] Correcting BUS records in tmp-kb-8f35dcc278fb4900b238db3d4c120f14/output.s.bus to tmp-kb-8f35dcc278fb4900b238db3d4c120f14/output.s.c.bus with on-list velocity/SRR6470906/10x_version2_whitelist.txt
[2025-10-29 10:58:24,943] INFO [count_lamanno] Sorting BUS file tmp-kb-8f35dcc278fb4900b238db3d4c120f14/output.s.c.bus to velocity/SRR6470906/output.unfiltered.bus
[2025-10-29 10:59:30,140] INFO [count_lamanno] Capturing records from BUS file velocity/SRR6470906/output.unfiltered.bus to tmp-kb-8f35dcc278fb4900b238db3d4c120f14/spliced.bus with capture list velocity/intron_t2c.txt
[2025-10-29 11:00:02,890] INFO [count_lamanno] Sorting BUS file tmp-kb-8f35dcc278fb4900b238db3d4c120f14/spliced.bus to velocity/SRR6470906/spliced.unfiltered.bus
[2025-10-29 11:00:23,621] INFO [count_lamanno] Inspecting BUS file velocity/SRR6470906/spliced.unfiltered.bus
[2025-10-29 11:00:26,426] INFO [count_lamanno] Generating count matrix velocity/SRR6470906/counts_unfiltered/spliced from BUS file velocity/SRR6470906/spliced.unfiltered.bus
[2025-10-29 11:00:58,495] INFO [count_lamanno] Capturing records from BUS file velocity/SRR6470906/output.unfiltered.bus to tmp-kb-8f35dcc278fb4900b238db3d4c120f14/unspliced.bus with capture list velocity/cdna_t2c.txt
[2025-10-29 11:01:13,821] INFO [count_lamanno] Sorting BUS file tmp-kb-8f35dcc278fb4900b238db3d4c120f14/unspliced.bus to velocity/SRR6470906/unspliced.unfiltered.bus
[2025-10-29 11:01:20,232] INFO [count_lamanno] Inspecting BUS file velocity/SRR6470906/unspliced.unfiltered.bus
[2025-10-29 11:01:22,136] INFO [count_lamanno] Generating count matrix velocity/SRR6470906/counts_unfiltered/unspliced from BUS file velocity/SRR6470906/unspliced.unfiltered.bus
[2025-10-29 11:01:41,457] INFO [count_lamanno] Reading matrix velocity/SRR6470906/counts_unfiltered/spliced.mtx
[2025-10-29 11:01:54,826] INFO [count_lamanno] Reading matrix velocity/SRR6470906/counts_unfiltered/unspliced.mtx
[2025-10-29 11:02:01,769] INFO [count_lamanno] Combining matrices
[2025-10-29 11:02:02,357] INFO [count_lamanno] Writing matrices to h5ad velocity/SRR6470906/counts_unfiltered/adata.h5ad
[2025-10-29 11:02:07,062] INFO [count_lamanno] Filtering with bustools
[2025-10-29 11:02:07,062] INFO [count_lamanno] Generating on-list velocity/SRR6470906/filter_barcodes.txt from BUS file velocity/SRR6470906/output.unfiltered.bus
[2025-10-29 11:02:08,466] INFO [count_lamanno] Correcting BUS records in velocity/SRR6470906/output.unfiltered.bus to tmp-kb-8f35dcc278fb4900b238db3d4c120f14/output.unfiltered.c.bus with on-list velocity/SRR6470906/filter_barcodes.txt
[2025-10-29 11:02:34,507] INFO [count_lamanno] Sorting BUS file tmp-kb-8f35dcc278fb4900b238db3d4c120f14/output.unfiltered.c.bus to velocity/SRR6470906/output.filtered.bus
[2025-10-29 11:02:59,446] INFO [count_lamanno] Capturing records from BUS file velocity/SRR6470906/output.filtered.bus to tmp-kb-8f35dcc278fb4900b238db3d4c120f14/spliced.bus with capture list velocity/intron_t2c.txt
[2025-10-29 11:03:28,192] INFO [count_lamanno] Sorting BUS file tmp-kb-8f35dcc278fb4900b238db3d4c120f14/spliced.bus to velocity/SRR6470906/spliced.filtered.bus
[2025-10-29 11:03:44,918] INFO [count_lamanno] Generating count matrix velocity/SRR6470906/counts_filtered/spliced from BUS file velocity/SRR6470906/spliced.filtered.bus
[2025-10-29 11:04:08,504] INFO [count_lamanno] Capturing records from BUS file velocity/SRR6470906/output.filtered.bus to tmp-kb-8f35dcc278fb4900b238db3d4c120f14/unspliced.bus with capture list velocity/cdna_t2c.txt
[2025-10-29 11:04:20,926] INFO [count_lamanno] Sorting BUS file tmp-kb-8f35dcc278fb4900b238db3d4c120f14/unspliced.bus to velocity/SRR6470906/unspliced.filtered.bus
[2025-10-29 11:04:27,037] INFO [count_lamanno] Generating count matrix velocity/SRR6470906/counts_filtered/unspliced from BUS file velocity/SRR6470906/unspliced.filtered.bus
[2025-10-29 11:04:46,158] INFO [count_lamanno] Reading matrix velocity/SRR6470906/counts_filtered/spliced.mtx
[2025-10-29 11:04:54,852] INFO [count_lamanno] Reading matrix velocity/SRR6470906/counts_filtered/unspliced.mtx
[2025-10-29 11:05:00,831] INFO [count_lamanno] Combining matrices
[2025-10-29 11:05:01,166] INFO [count_lamanno] Writing matrices to h5ad velocity/SRR6470906/counts_filtered/adata.h5ad
✓ count workflow completed!
dict_keys(['workflow', 'technology', 'output_path', 'parameters'])
Analysis¶
In this part of the tutorial, we will load the RNA count matrix generated by kb count into Python and analyse them using the omicverse pipeline.
You can find detailed information about these codes in this website.
adata = sc.read_h5ad("velocity/SRR6470906/counts_filtered/adata.h5ad")
adata.layers['spliced'] = adata.layers['mature']
adata.layers['unspliced'] = adata.layers['nascent']
velo_obj=ov.single.Velo(adata)
velo_obj.filter_genes(min_shared_counts=20)
velo_obj.preprocess()
In Velo module, you should keep all genes' expression not normalized.
Filtered out 53587 genes that are detected 20 counts (shared).
|-----> Running monocle preprocessing pipeline...
|-----> convert ensemble name to official gene name
|-----? Your adata object uses non-official gene names as gene index.
Dynamo is converting those names to official gene names.
|-----> Storing myGene name info into local cache db: mygene_cache.sqlite.
|-----> Subsetting adata object and removing Nan columns from adata when converting gene names.
|-----------> filtered out 1151 outlier cells
|-----------> filtered out 1502 outlier genes
|-----> PCA dimension reduction
|-----> <insert> X_pca to obsm in AnnData Object.
|-----> computing cell phase...
|-----> [Cell Phase Estimation] completed [65.5472s]
|-----> [Cell Cycle Scores Estimation] completed [0.3818s]
|-----> [Preprocessor-monocle] completed [13.5334s]
🖥️ Using Scanpy CPU to calculate neighbors...
🔍 K-Nearest Neighbors Graph Construction:
Mode: cpu
Neighbors: 30
Method: umap
Metric: euclidean
Representation: X_pca
PCs used: 30
computing neighbors
🔍 Computing neighbor distances...
🔍 Computing connectivity matrix...
💡 Using UMAP-style connectivity
✓ Graph is fully connected
✅ KNN Graph Construction Completed Successfully!
✓ Processed: 4,331 cells with 30 neighbors each
✓ Results added to AnnData object:
• 'neighbors': Neighbors metadata (adata.uns)
• 'distances': Distance matrix (adata.obsp)
• 'connectivities': Connectivity matrix (adata.obsp)
finished: added to `.uns['neighbors']`
`.obsp['distances']`, distances for each pair of neighbors
`.obsp['connectivities']`, weighted adjacency matrix (0:00:00)
velo_obj.moments(backend='scvelo')
velo_obj.dynamics(backend='scvelo',n_jobs=8)
velo_obj.cal_velocity(method='scvelo')
WARNING: Did not normalize X as it looks processed already. To enforce normalization, set `enforce=True`.
Normalized count data: spliced, unspliced.
computing moments based on connectivities
finished (0:00:05) --> added
'Ms' and 'Mu', moments of un/spliced abundances (adata.layers)
recovering dynamics (using 8/384 cores)
finished (0:01:22) --> added
'fit_pars', fitted parameters for splicing dynamics (adata.var)
computing velocities
finished (0:00:02) --> added
'velocity', velocity vectors for each individual cell (adata.layers)
velo_obj.velocity_graph(
vkey='velocity_S',
xkey='Ms',
n_jobs=8,
)
computing velocity graph (using 8/384 cores)
finished (0:00:03) --> added
'velocity_S_graph', sparse matrix with cosine correlations (adata.uns)
ov.pp.neighbors(adata, n_neighbors=15,
use_rep='X_pca')
ov.pp.umap(adata)
ov.pp.leiden(adata,resolution=0.2)
🖥️ Using Scanpy CPU to calculate neighbors...
🔍 K-Nearest Neighbors Graph Construction:
Mode: cpu
Neighbors: 15
Method: umap
Metric: euclidean
Representation: X_pca
computing neighbors
🔍 Computing neighbor distances...
🔍 Computing connectivity matrix...
💡 Using UMAP-style connectivity
✓ Graph is fully connected
✅ KNN Graph Construction Completed Successfully!
✓ Processed: 4,331 cells with 15 neighbors each
✓ Results added to AnnData object:
• 'neighbors': Neighbors metadata (adata.uns)
• 'distances': Distance matrix (adata.obsp)
• 'connectivities': Connectivity matrix (adata.obsp)
finished: added to `.uns['neighbors']`
`.obsp['distances']`, distances for each pair of neighbors
`.obsp['connectivities']`, weighted adjacency matrix (0:00:00)
🔍 [2025-10-29 11:08:02] Running UMAP in 'cpu' mode...
🖥️ Using Scanpy CPU UMAP...
🔍 UMAP Dimensionality Reduction:
Mode: cpu
Method: umap
Components: 2
Min distance: 0.5
{'n_neighbors': 15, 'method': 'umap', 'random_state': 0, 'metric': 'euclidean', 'use_rep': 'X_pca'}
🔍 Computing UMAP parameters...
🔍 Computing UMAP embedding (classic method)...
✅ UMAP Dimensionality Reduction Completed Successfully!
✓ Embedding shape: 4,331 cells × 2 dimensions
✓ Results added to AnnData object:
• 'X_umap': UMAP coordinates (adata.obsm)
• 'umap': UMAP parameters (adata.uns)
✅ UMAP completed successfully.
🖥️ Using Scanpy CPU Leiden...
running Leiden clustering
finished: found 12 clusters and added
'leiden', the cluster labels (adata.obs, categorical) (0:00:00)
velo_obj.velocity_embedding(
basis='umap',
vkey='velocity_S',
)
computing velocity embedding
finished (0:00:00) --> added
'velocity_S_umap', embedded velocity vectors (adata.obsm)
fig = ov.plt.figure(figsize=(4, 4))
ax = ov.plt.subplot(1, 1, 1)
ov.pl.embedding(
adata,
basis='X_umap',
color='leiden',
ax=ax,
show=False,
size=50,
alpha=0.3
)
ov.pl.add_streamplot(
adata,
basis='X_umap',
velocity_key='velocity_S_umap',
ax=ax,
)
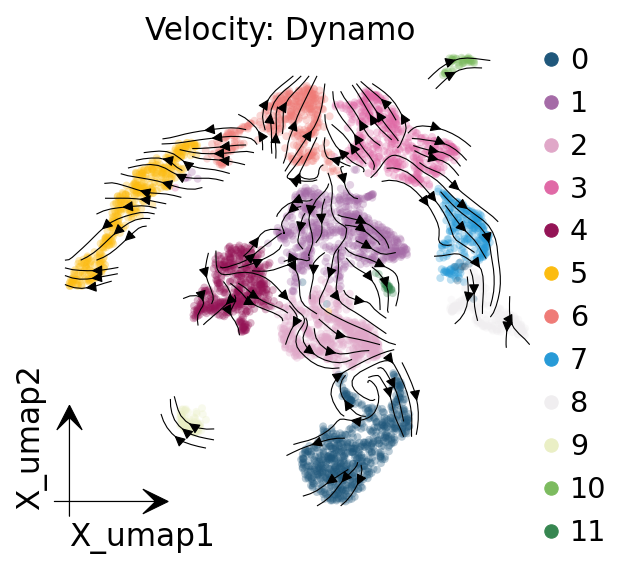
ov.plt.title('Velocity: Dynamo')