Using OmicVerse MCP with Claude Code — Step by Step¶
This page focuses on the Claude Code workflow: how to connect OmicVerse MCP, verify the tools, run a standard analysis, and inspect the results. Broader deployment details, the full tool catalog, and runtime internals now live in the dedicated MCP pages.
Related Pages¶
- Complete onboarding: Full Start
- Deployment patterns: Clients and Deployment
- Full tool inventory: Tool Catalog
- Runtime behavior and troubleshooting: Runtime and Troubleshooting
- Flags and JSON-RPC reference: Reference
Minimal Claude Code Setup¶
Project-level .mcp.json with stdio¶
{
"mcpServers": {
"omicverse-local": {
"type": "stdio",
"command": "python",
"args": [
"-m", "omicverse.mcp",
"--phase", "P0+P0.5",
"--persist-dir", "/tmp/ov_persist_local"
],
"env": {}
}
}
}
Global shortcut¶
claude mcp add omicverse -- python -m omicverse.mcp --phase P0+P0.5
Local HTTP option¶
Start OmicVerse MCP:
NUMBA_CACHE_DIR=/tmp/numba_cache MPLCONFIGDIR=/tmp/mpl \
python -m omicverse.mcp \
--transport streamable-http \
--host 127.0.0.1 \
--port 8765 \
--http-path /mcp \
--phase P0+P0.5
Then point Claude Code at it:
{
"mcpServers": {
"omicverse": {
"type": "http",
"url": "http://127.0.0.1:8765/mcp"
}
}
}
Step-by-Step Walkthrough¶
Step 1: Launch Claude Code¶
Navigate to your project directory and start Claude Code:
cd /path/to/your/project
claude
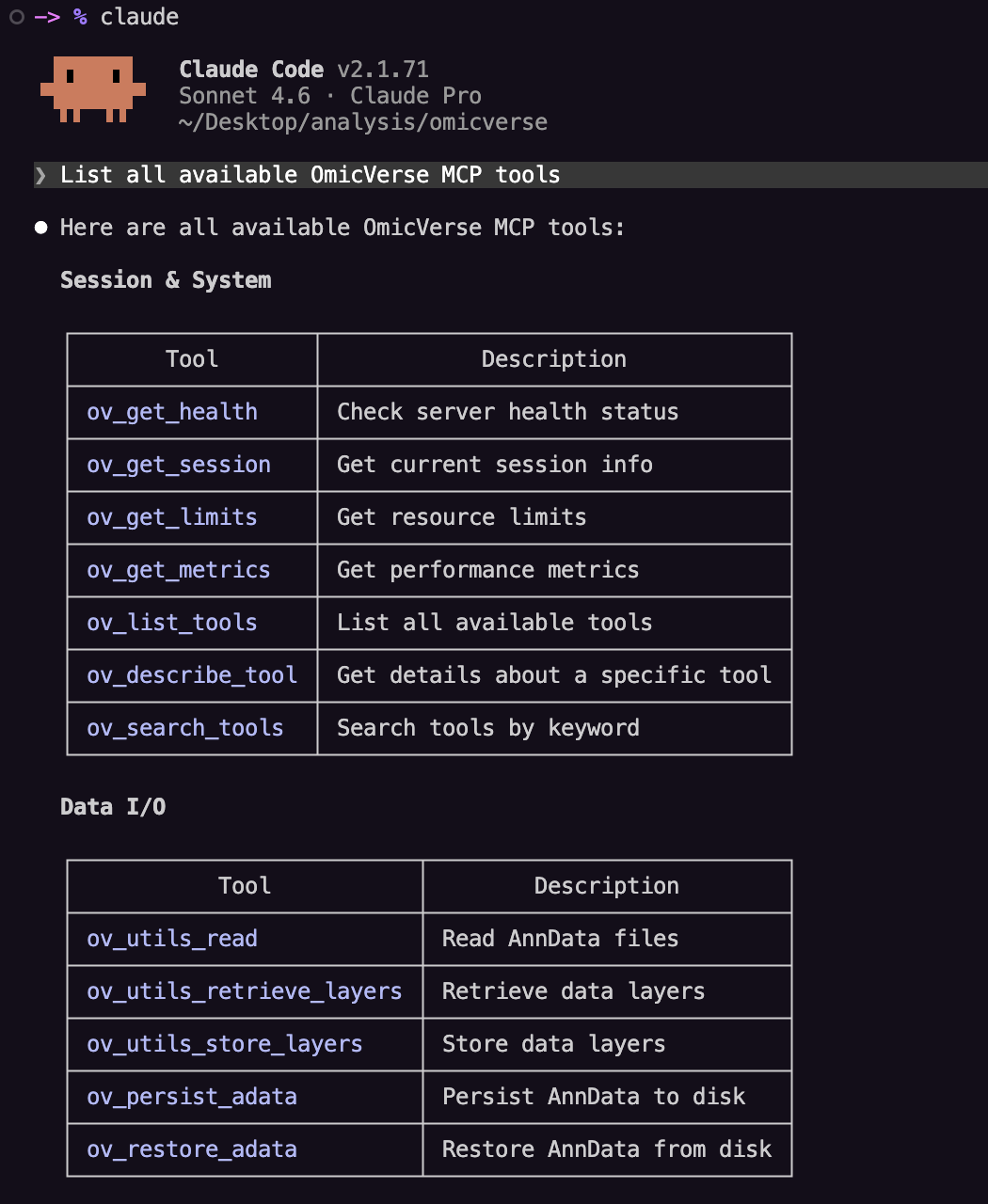
Step 2: Verify tools are loaded¶
Ask:
List all available OmicVerse MCP tools
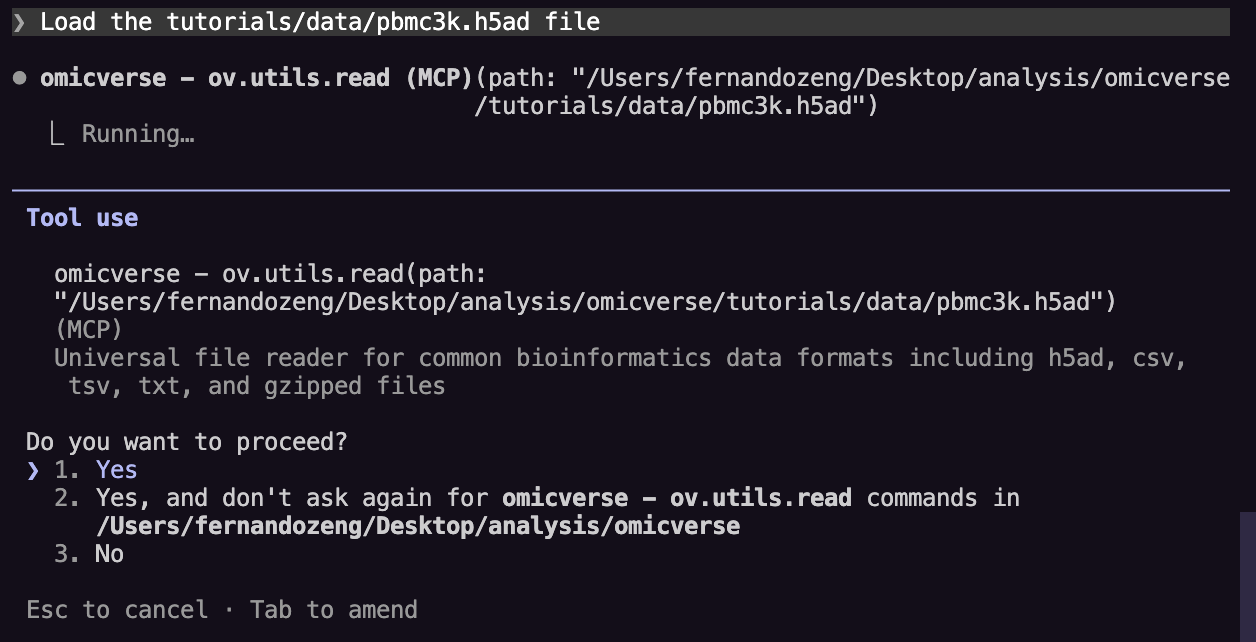
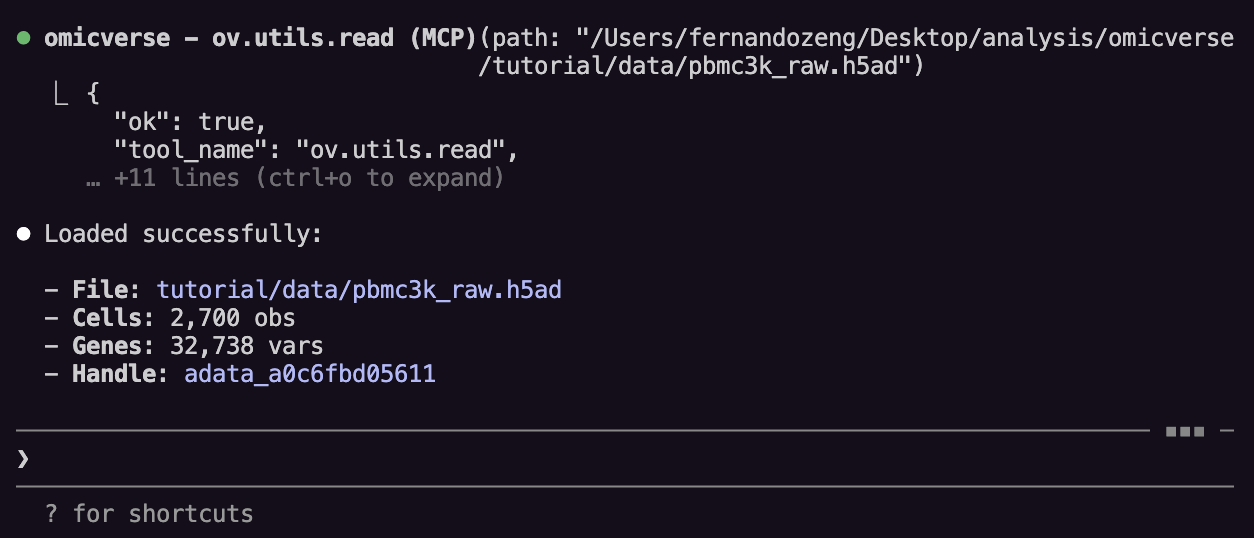
Step 3: Load your data¶
Ask:
Load the pbmc3k.h5ad file
Or use a built-in dataset:
Load the built-in seqfish dataset
The adata_id is a server-side reference. Claude tracks it automatically.
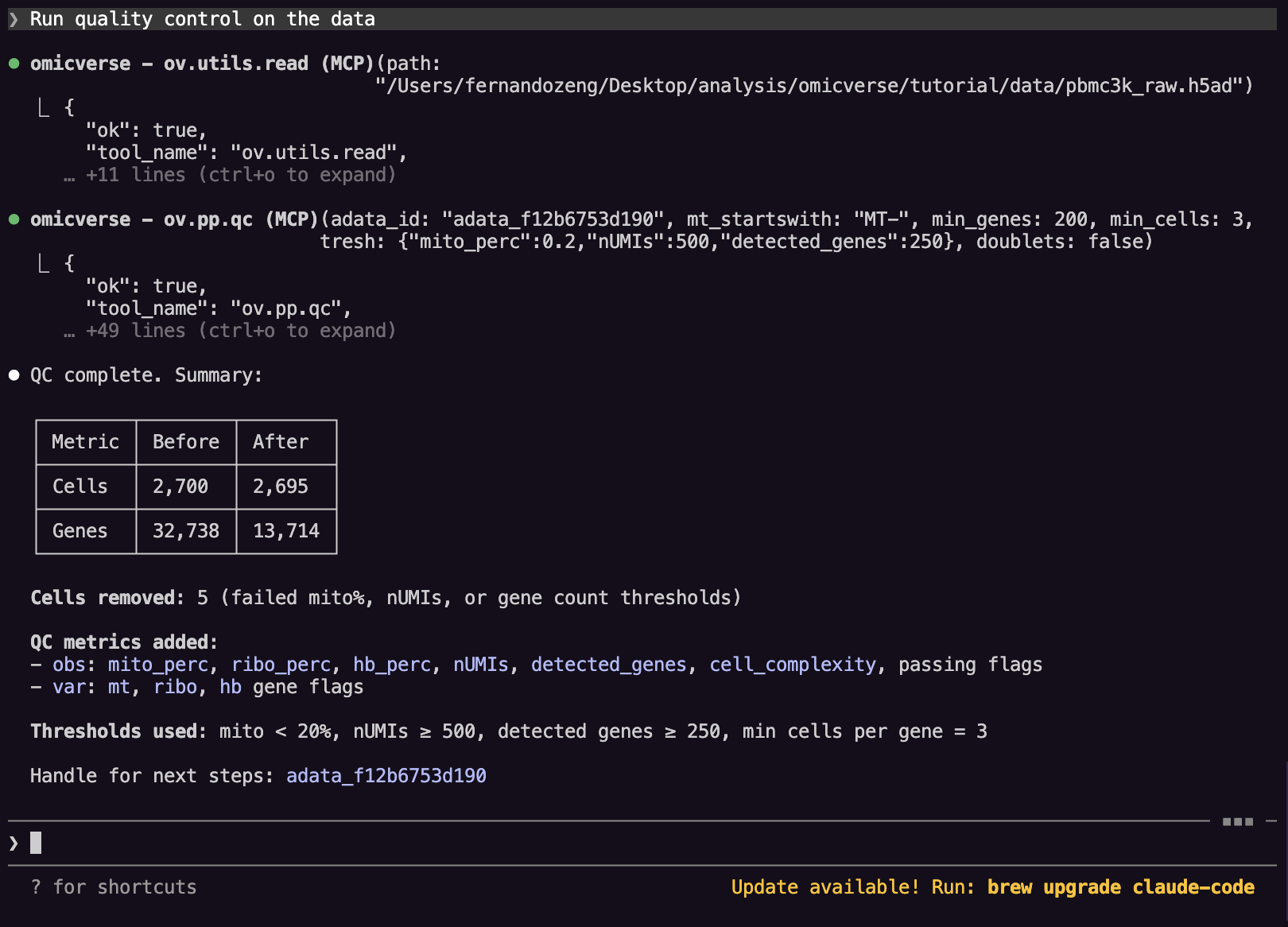
Step 4: Quality control¶
Ask:
Run quality control on the data
Claude calls ov.pp.qc, which computes per-cell metrics such as:
n_genesn_countspct_counts_mt
Step 5: Run preprocessing¶
Ask:
Run the standard preprocessing: scale, PCA with 50 components, build neighbors, compute UMAP, and do Leiden clustering at resolution 1.0
Claude runs the tools in sequence:
ov.pp.scaleov.pp.pcaov.pp.neighborsov.pp.umapov.pp.leiden
Automatic prerequisite checking¶
The MCP server enforces the correct ordering. If Claude tries to skip a step, the server returns an error such as:
{"ok": false, "error_code": "missing_data_requirements", "suggested_next_tools": ["ov.pp.scale"]}
Claude can use suggested_next_tools to self-correct.
Step 6: Visualize¶
Ask:
Plot the UMAP colored by Leiden clusters
Show me a violin plot of n_genes grouped by leiden cluster
Create a dot plot for genes CD3D, CD79A, LYZ, NKG7 grouped by leiden
Step 7: Marker analysis¶
Ask:
Find marker genes for each Leiden cluster and show me the top 5 per cluster
Plot a marker gene dotplot
Run COSG to rank marker genes
Perform pathway enrichment on the marker genes
Plot the pathway enrichment results
Step 8: Save and restore¶
Ask:
Save the current dataset to disk
Later:
Restore the dataset from /path/to/file.h5ad
Complete Conversation Example¶
You: Load the file pbmc3k.h5ad
Claude: Loaded AnnData with 2700 cells x 32738 genes.
You: Run QC, preprocessing, and cluster the cells
Claude: I'll run the full pipeline step by step.
[calls ov.pp.qc]
[calls ov.pp.scale]
[calls ov.pp.pca]
[calls ov.pp.neighbors]
[calls ov.pp.umap]
[calls ov.pp.leiden]
You: Show me the UMAP
Claude: [calls ov.pl.embedding]
You: Find marker genes and show top 3 per cluster
Claude: [calls ov.single.find_markers]
[calls ov.single.get_markers]
You: Save the results
Claude: [calls ov.persist_adata]
P2 Workflows¶
If you need advanced class-backed tools, start OmicVerse with:
{
"mcpServers": {
"omicverse": {
"command": "python",
"args": ["-m", "omicverse.mcp", "--phase", "P0+P0.5+P2"]
}
}
}
This enables workflows such as:
ov.bulk.pydegov.single.pyscsaov.single.metacellov.single.dctov.utils.lda_topic
pyDEG¶
Typical lifecycle:
ov.bulk.pydegwithcreateov.bulk.pydegwithrunov.bulk.pydegwithresultsov.bulk.pydegwithdestroy
pySCSA¶
Typical lifecycle:
ov.single.pyscsawithcreateov.single.pyscsawithannotateov.single.pyscsawithdestroy
MetaCell¶
Typical lifecycle:
ov.single.metacellwithcreateov.single.metacellwithtrainov.single.metacellwithpredictov.single.metacellwithdestroy
Short Troubleshooting List¶
- Missing tools: ask Claude to run
ov.list_tools - Missing dependencies: ask Claude to run
ov.describe_tool - Lost session state: reload or
ov.restore_adata - Long-running task confusion: inspect Runtime and Troubleshooting