OmicVerse Documentation¶
A single pipeline for the entire transcriptome universe¶
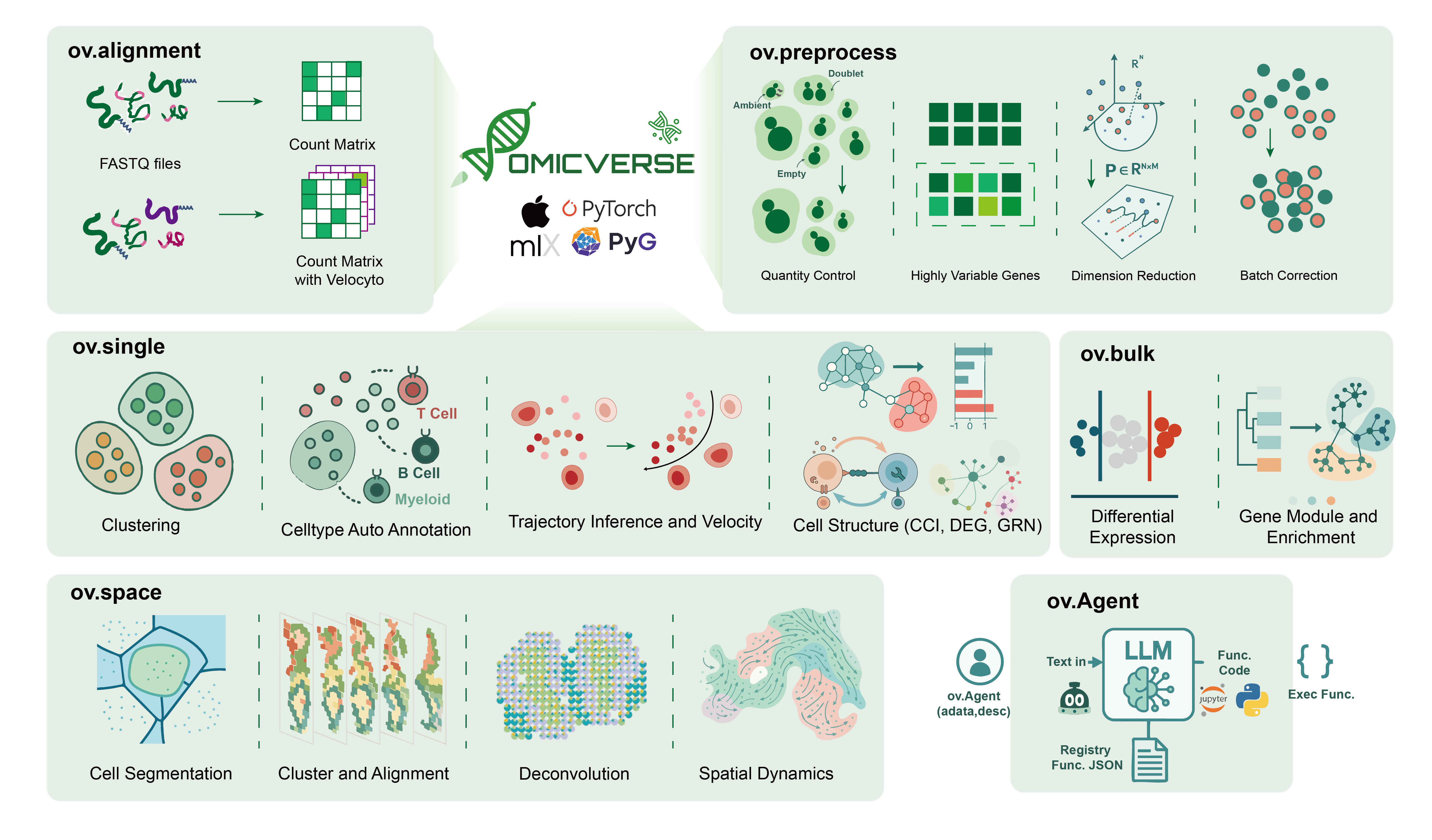
OmicVerse v2 is a unified Python framework for modern transcriptomics and multi-omics analysis. It brings together bulk RNA-seq, single-cell, spatial transcriptomics, visualization, model-based analysis, and AI-assisted workflows in one package.
If you find OmicVerse useful for your research, please consider citing the OmicVerse manuscript.
New to omicverse? Set up your environment with conda, pip, or Docker.
Step-by-step tutorials covering bulk, single-cell, spatial, and AI-assisted workflows.
Detailed description of every public function and class in OmicVerse.
Deploy OmicVerse as a conversational AI agent via Telegram, Feishu, MCP, and more.
Need help? Reach out on our GitHub Discussions forum.
Find a bug? Interested in contributing? Check out our GitHub.